Overview
Instinctive behaviors, such as feeding, escape and mating, are an animal’s survival kit of actions that they acquire with little to no experience. Although instinctive behaviors are generated by remarkably conserved brain circuits, it has become increasingly clear in recent years that instinctive behaviors are not just stereotyped fixed-action patterns but that they can be flexible in regard to both action selection and execution. They are strongly modulated by internal and motivational states, such as stress, hunger, emotion (fear/anxiety), and/or circadian cycle, and may undergo short- and long-lasting adaptations to accommodate environmental changes.
Understanding the synaptic and cellular implementation of this behavioral flexibility in defined neural circuits will be a significant step towards our understanding of adaptive information processing in the brain and the generation of behavior. We will focus on evolutionarily conserved nociceptive circuits in insects and rodents, that are critically involved in the initiation and execution of instinctive behaviors such as escape from noxious environments. Our goal is to identify canonical neuronal computations and plasticity mechanisms that impart flexibility to instinctive behaviors.
To address these issues, we use a multidisciplinary approach. We perform in vivo neural activity recordings and manipulation experiments in ethologically-relevant behavioral tasks in fly and mice, and combine these with molecular, cellular and circuit-level analyses in vitro.
<Recent publications>
1. Molecular and cellular control of developmental neural plasticity
・Furusawa et al. Science (2023).
・Togashi et al. Front Cell Neurosci (2020).
・Kitatani et al. PLoS Genetics (2020).
・Yasunaga et al. Genes & Development (2015).
・Kanamori et al. Nature Communications (2015).
・Kanamori et al. Science (2013).
2. Flexible control of sensory processing and behavior
・Nakamizo-Dojo et al. Nature Communications (2023).
・Tsuji et al. Nature Communications (2023).
・Takeuchi et al. Communications Biology (2023).
・Nakahama et al. Molecular Brain (2022).
・Omamiuda-Ishikawa et al. PLoS Genetics (2020).
・Utashiro et al. Scientific Reports (2018).
・Yoshino et al. Current Biology (2017).
<class=”mb20″ span=”” style=”color: blue;” color=”blue”>1. Receptive field determination in sensory neurons</class=”mb20″>
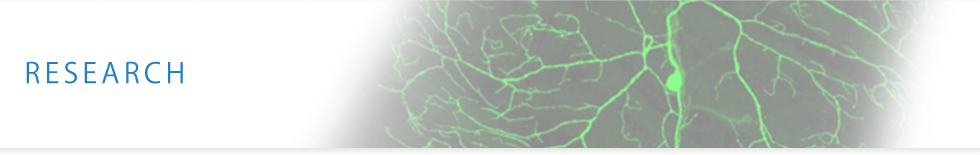
 The third instar larvae with GFP reporter in a subset of sensory neurons (left panel). Dendritic tiling of sensory neurons in wild-type and trc mutants (Emoto et al. Cell 2004).
The third instar larvae with GFP reporter in a subset of sensory neurons (left panel). Dendritic tiling of sensory neurons in wild-type and trc mutants (Emoto et al. Cell 2004).For precise sampling of sensory information, certain neurons arrange tiling of their dendrites to cover the receptive field completely and non-redundantly. By using fruit fly sensory neurons as a model system, we have shown that the dendritic tiling is mediated by homotypic repulsion between neighboring dendrites that the repulsive interaction is mediated by the NDR kinase signaling pathway (Cell 2004; Nature 2006; Neuron 2007; EMBO 2009). We are currently conducting a genome-wide genetic screen in Drosophila sensory neurons to identify the whole molecular network underlying the dendritic tiling.
【References】
・Emoto et al. Cell 2004
・Emoto et al. Nature 2006
・Soba et al. Neuron 2007
・Parrish et al. Genes Dev. 2007
・Koike-Kumagai et al. EMBO J. 2009
<class=”mb20″ span=”” style=”color: blue;” color=”blue”>2. Structural plasticity of neuronal circuits</class=”mb20″>
Neuronal circuits in the brain are not static. In many systems, especially during critical periods of development, neurons exhibit a period of juvenile plasticity in which connectivity can be modified in response to sensory input or following specific experiences, thereby providing neurons with new response properties, tailored to the new environment. To achieve these changes in connectivity, certain neurons modify the shape of their axon and dendritic arbors in response to various stimuli. We have identifies novel mechanisms that regulate structural plasticity of dendritic arbors in fly sensory neurons (Genes Dev. 2007; Dev. Cell 2010). We are investigating how these mechanisms are related to the sensory-evoked plasticity in the fly and mice nervous system.
【References】
・Togashi et al. Frontiers in Cell Neuroscience 2020
・Kitatani et al. PLoS Genetics 2020
・Kanamori et al. Nature Communications 2015
・Yasunaga et al. Dev. Cell 2010
・Emoto, K. Curr. Opin. Neurobiol. 2012
・Kanamori et al. Science 2013
<class=”mb20″ span=”” style=”color: blue;” color=”blue”>3. Neuronal mechanisms underlying preference change</class=”mb20″>
The behavior of animals is flexible and can change dramatically based on the environment, nutritional state, or even age, among other factors. However, the neural basis of how external and internal cues modify innate behavior is not clearly understood. Appropriate preferences for light or dark conditions can be critical for an animal’s survival. Innate light preferences are not static in some animals, including the fruit fly, which prefers darkness in the feeding larval stage but prefers light in adulthood. This suggests that the neuronal circuits that determine the light preference have been modified during metamorphosis. To elucidate the neuronal mechanisms underlying the preference change, we examine neurons involved in larval phototactic behavior by regulating neuronal functions. To do that, we have established a quantitative assay system for larval phototaxis and started genetic screens. Through this study, we would like to understand how the brain makes decisions and also change the decision against the same sensory information.
【References】
・Omamiuda-Ishikawa et al. PLoS Genetics (2020)
・Yoshino et al. Curr Biol (2017)
・Morikawa et al. PNAS (2011)
・Sakurai et al. Nature Communications (2013)
<class=”mb20″ span=”” style=”color: blue;” color=”blue”>4. Adult neurogenesis in the brain</class=”mb20″>
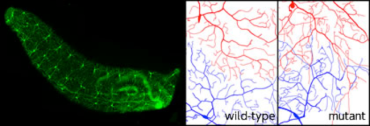
 Newly generated neurons in the subventricular zone (SVZ) migrate into olfactory bulb (Left) (Koizumi et al. Nat. Neurosci. 2006). Neural stem cells (neuroblasts) in the third instar larval brain.
Newly generated neurons in the subventricular zone (SVZ) migrate into olfactory bulb (Left) (Koizumi et al. Nat. Neurosci. 2006). Neural stem cells (neuroblasts) in the third instar larval brain.Neural stem cells have been found in certain areas in the adult brain. Although the adult neurogeneis likely plays an important role in learning and memory, the information is still limited. We are investigating the roles of newborn neurons in the mouse and fruit fly brain.
【References】
・Koizumi et al. Dev Neurobiol 2017
・Koizumi et al. Neuron 2006c
・Koizumi et al. Nature Neurosci. 2006
<class=”mb20″ span=”” style=”color: blue;” color=”blue”>5. “do we run coz we’re afraid, or are we afraid coz we run?”
“does our heart race coz we’re afraid, or are we afraid coz our heart race?” </class=”mb20″>
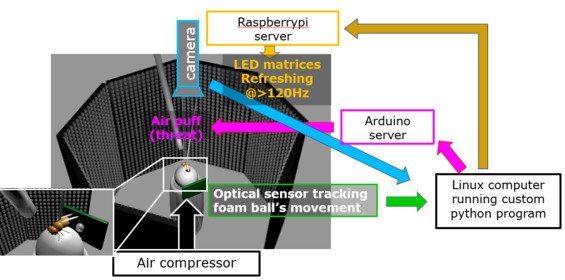 Custom-built virtual reality paradigm for flies
Custom-built virtual reality paradigm for fliesFrightened animals run (escape), increase the heart rate (heart regulation), and exaggerate surprise by faint sound (cognitive bias), in a coordinated fashion. There has been a long dispute in the field of phylosophy and psychology as to “do we run coz we’re afraid, or are we afraid coz we run?” “does our heart race coz we’re afraid, or are we afraid coz our heart race?”. We embarked on tackling this fundamental problem, which has so far been examined philosophically/psychologically, by newly bringing neurogenetical methodologies. We have been the first to establish a novel Drosophila model suitable for tackling above problem, by discovering that even Drosophila when threatened shows escape, heart regulation, and cognitive bias. Currently we are rushing to identify molecular/neuronal substrates responsible for these suit of chages and to reveal fairly characteristic activity pattern of such neurons.
○Key words○
componential theory of fear
escape
heart regulation
cognitive bias
philosophy/psychology meet neurogenetics
synchronized firing
virtual reality paradigm
Ca imaging of brain
<class=”mb20″ span=”” style=”color: blue;” color=”blue”>6. Neural basis for human intelligence</class=”mb20″>
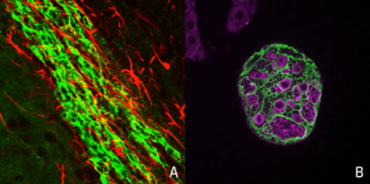
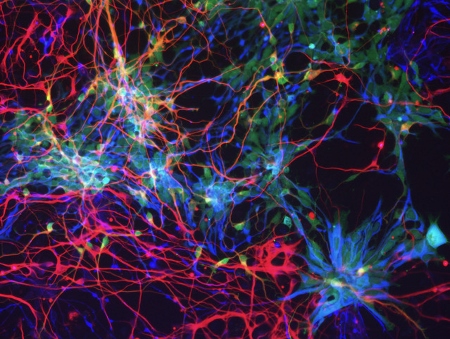 The cortical cells differentiated from human ES cell in vitro. The morphology of the neural stem cells (blue) and neurons (red) are visualized by GFP expression (green).
The cortical cells differentiated from human ES cell in vitro. The morphology of the neural stem cells (blue) and neurons (red) are visualized by GFP expression (green).Our species, human, is characterized by the higher cognitive functions such as the language-based communication and the conceptual thinking, Most of these functions are elicited by the computational power of the cerebral cortex. Human has significantly larger volume of cerebral cortex than the evolutionary cousins such as the chimpanzee and the gorilla, and such an extraordinarily expanded cortex facilitates our cognitive abilities. Recent advancement of genomics allowed us to identify a repertoire of genes specifically detected in the human genome but not in those of other species, most of which are originated by recent gene duplication. Some of these “human-specific genes” are actually involved in the brain development and function. For instances, we found that one of human-specific gene called NOTCH2NL has a pivotal role in expanding neuronal production and consequently the enlargement of cortical volume. Human-specific genes are also important to understand the disease mechanisms in addition to the evolution, because a significant fraction of genomic regions encoding human-specific genes including NOTCH2NL are reported to be linked to congenital neurodevelopmental and cognitive diseases. We aim to reveal the genetic and molecular mechanisms of human brain evolution by the comparative genomics and the experimental approaches based on in vitro models of human brain development using the pluripotent stem cells such as ES cell and iPS cell at the level of (1) genome, (2) cell, (3) brain structure and neural circuit embedded in it, and (4) behaviour and the disease phenotypes.
【References】
・Suzuki et al Developmental Cell 22 863 (2012)
・Suzuki et al Cell 173 1370 (2018)
Fight or flight: genes and neurons controlling animal aggression.
Animals are continuously exposed to competition with each other for limited resource in their natural environment. The confronting individuals go through a series of decision-making processes on whether to straightly fight or to retreat in avoidance of further risk of damage. However, molecular mechanisms underlying such behavioral switching between “fight” or “flight” remain elusive. In our group, we address this issue by exploiting reagents that provide genetic and neuronal control of animal aggression.
Recent studies by us and others have identified several molecules and neurons important for spontaneous execution of aggression by using fruit flies (Drosophila melanogaster) as a genetically tractable model. These findings, however, brought us to the next key question: how the attacked individuals respond to their opponents. With the combined power of fly genetics and machine-learning-based behavioral analyses, we aim to mechanistically elucidate the following processes: (i) physical/non-physical reception of attacks/threats, (ii) context-dependent value judgement on whether to make a counterattack, and (iii) subsequent behavioral transitions. Such decision-making directly affects the outcome of each battle and eventually leads to hierarchical relationship within a population, underscoring its significance in various forms of animal societies.
Yet another aspect of aggression is the related neuropathologies in humans such as those observed in post-traumatic stress disorder (PTSD). Interestingly, studies with mammalian PTSD models have recently suggested the role of innate immunity in altering neuronal functions under chronically stressed conditions. For the evolutionary conservativity of innate immunity, the fruit fly has a potential to provide a novel approach to molecularly unravel how such system developed for combat with internal microbes could also modulate the fight towards external enemies.
○Key words○
Aggression
Escape
Decision-making
Nociception
Optogenetics
Machine-learning
Hierarchical relationship
Social interactions
PTSD
Innate immunity
【References】
・Ishii K*, Wohl M*, DeSouza A, Asahina K. [*Equal contribution] (2020) eLife
・Wohl M*, Ishii K*, Asahina K. [*Equal contribution] (2020) eLife
・Leng X, Wohl M, Ishii K, Nayak P, Asahina K. (2020) PLoS One
・Ishii K, Cortese M, Leng M, Shokhirev MN, Asahina K. (2020) bioRxiv
・Nie X, et al. (2018) Neuron
・Ishii K, Hamamoto H, Sekimizu K. (2015) Arch Insect Biochem Physiol